Mycorrhizas (4)
The Ectomycorrhiza Community:
Uncovering the Foundation of Our Temperate Forests
Walk through a temperate or boreal (far northern) forest or woodland at the right time of year and you'll find mushrooms produced by ectomycorrhizal fungi, sometimes quite a lot of them. Do this often enough and in enough different places, you'll find that different types of mushrooms occur in different types of forest. A Scots pine (Pinus sylvestris) woodland in Britain or Sweden can produce Suillus in abundance, yet invariably they will be absent from a nearby Norway spruce (Picea abies) plantation. Lactarius rufus and Amanita muscaria, on the other hand, might well be present in both. Ecologists refer to the collection of organisms in a particular area or habitat as a community. Thus, each root, tree, and woodland has its own community of mycorrhizal fungi and they comprise part of the overall biological community of that tree or woodland.
Although knowing which fungi make up the ectomycorrhiza community of an area is interesting in itself, more importantly it is the first step in learning how this important part of the forest works. We know that ectomycorrhizal fungi can play many roles in a forest -- for instance, increasing the availability and plant uptake of a variety of nutrients such as nitrogen, phosphorus, and potassium, protecting plant roots from attack by pathogenic fungi, and increasing drought resistance. However, we know very little about the details of who's doing what, fungus-wise.
Although our knowledge of the particular talents of different fungi is still rudimentary, it is clear that there is considerable diversity among them with respect to their functions. When you've seen one, you haven't seen them all; or, in the jargon of ecologists, there is much less functional redundancy among ectomycorrhizal fungi than has usually been thought.
But why are there so many different ectomycorrhizal fungi? Does an individual tree need all of the many fungi that can occur simultaneously on its roots? Does one fungus provide nitrogen, while another provides phosphorus, while another guards against attack by potential root rotters? How much control does a tree have over which fungi develop mycorrhizas with its roots? What happens to the trees if inputs of fertilizer, or acid rain, or climate change cause some fungi to decline or disappear, while others become more abundant?
Ultimately, the answers to these and many other important questions will hinge on our ability to figure out the physiological functioning of different fungus species, in association with different plants, in different environments. But before we can begin such studies, we have to know who is associating with whom, how many, and when, and where. In short, we need to understand community composition and structure.
The first step in studying communities is to identify their constituents. Three main approaches for identifying ectomycorrhizal fungi in forests have been followed:
- Collect, describe, and identify the mushrooms produced by ectomycorrhizal fungi (the main approach through the mid 1990's).
- Collect, describe, and identify, based on their outward appearance (morphology) and inner structure (anatomy), the mycorrhizal root tips in the soil (this is referred to as "morphotyping"; it became well established in the mid- to late 1980's).
- Collect, describe, and identify, based on molecular characteristics (of DNA, proteins, and other complex biological chemicals), the mycorrhizal root tips (this began in the early 1990's and expanded rapidly in the mid-90's).
We'll begin by taking a look at each of these approaches to see how they work and what limitations they have. Then we'll review some of the research findings to see what these methods are telling us about the ectomycorrhizal community at particular forest sites.
Mushroom Surveys (above-ground studies)
Until fairly recently, collecting mushrooms was the only means we had of establishing what species of fungi made up the ectomycorrhiza community of a site. It is still the simplest, least expensive, and most straightforward method. However, this approach has obvious limitations, most of which will be frustratingly familiar to every mushroom hunter. For instance becoming a proficient mushroom identifier takes a long time. In addition, mushrooms don't fruit predictably each year, as both anecdotal evidence and careful record-keeping have shown.
In the book, Mr. Jackson's Mushrooms, an excerpt from the notebooks of Canadian naturalist and artist Henry A.C. Jackson (for whom the 1984 NAMA Foray and Amanita jacksonii, the beautiful orange "Caesar's mushroom" of eastern North America, were named) recounts how, while walking in the woods on an August day in 1950, he "let out a whoop when I recognized it (a blue-black clustered chanterelle) as Cantharellus multiplex (now called Polyozellus multiplex) which I found at St. Come, Québec, fifteen years ago and which created so much interest and brought me recognition." His find in 1935 had been the first record of this fungus in Canada and apparently he had not found it since.
In a well documented example reported by R.J. Tofts and P.D. Orton in the August 1998 issue of Mycologist, 21 years of collecting in a Scots pine woodland in England yielded interesting data on mushroom fruiting. The principal objective of the project was to determine the number of years of collecting needed to establish a reasonably complete species list for the site (the authors wisely noted that a fully complete list was an unattainable ideal). They found that, even after 21 years, each new season of collecting produced, on average, 15 to 20 new species for the list. Among their other findings:
- Even though the overall species list continued to rise, the list of species that are most characteristic of Scots pine woods did show signs of leveling off.
- The overwhelming majority of species are uncommon -- approximately one-third were recorded only once, less than 12% were recorded in at least half the years, and none were recorded in all 21 years.
Two important consequences of these findings were pointed out by Tofts and Orton. First, it is clear that species lists (and, thus, our notions about the community) for different years of collecting would be different. Second, collecting over 21 years was not enough to compile a reasonably complete list for the site. The authors suggested that at least 25 to 30 years would be necessary.
Thus, collecting mushrooms over a 1- to 3-year period, the typical duration of most academic research projects, can give only a minimal picture of the species comprising the ectomycorrhiza community, and yields little, if any, information on their relative abundance on roots or as hyphae in the soil. In addition, because most woodlands are made up of more than one tree species, careful recording of which trees are growing near which mushrooms and comparison of many such records is necessary to provide us with an idea of which fungus species are associating with which tree species in ectomycorrhizas. Thus, logic suggests that it might be more productive to shift our focus below ground, and look directly at the roots.
Morphotyping
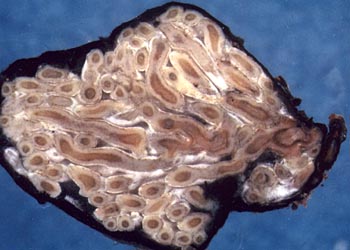
Different combinations of plants and fungi produce different-looking ectomycorrhizas (Figure 1). Thus, mycorrhizal root tips can be removed from soil samples (often with a great deal of painstaking work using needles, brushes, and fine forceps under a dissecting microscope) and described, in much the same way mycologists describe in their notes the characteristics of the mushrooms they collect and deposit in herbariums. A wide range of characters has been used to describe ectomycorrhizas ("morphotypes"), and there is no consensus as to which are most informative and how many must be recorded for a minimally useful description. Two growing collections of well illustrated descriptions have been published to date -- a fairly large one, primarily for Europe, and a more recent smaller one for North America (mostly the Pacific Northwest). However, these collections cover only a small percentage of the total species and, in many cases, the actual identity of the fungus is not known, so provisional names must be assigned to the mycorrhizas. So, why is the morphotype information so sparse?
 |
 |
 |
 |
 |
 |
Figure 1: Examples of ectomycorrhiza morphotypes. Different combinations of fungi and plants produce different-looking mycorrhizas. (a) Hebeloma crustuliniforme and Douglas-fir. (b) Cortinarius sp. and Douglas-fir. (c) Melanogaster intermedius and Pacific madrone. (d) Amanita muscaria and Sitka spruce. (e) Rhizopogon vinicolor and Douglas-fir. (f) Lactarius rubrilacteus and Douglas-fir. The different photos encompass total widths from about 1 (e) to 3.5 (a and f) centimeters.
Photos by Randy Molina (a, c, d), B. Zak (b, f), and Dan Luoma (e); used with permission. Thanks to Tom Horton for providing copies of the photos, which appeared previously in Horton and Bruns (2001).
|
|
First, to clearly observe many of the important characteristics, a high-quality compound microscope with special optical systems is needed (such as differential interference contrast, commonly referred to as Nomarski optics), and such scopes can easily cost more than $25,000. Second, even with a top-notch scope, the process of making a good description is terribly tedious and very time-consuming, and few persons have the combination of time, skill, patience, and funding to do it. Third, and perhaps most importantly, having a description is of little use until the identity of the fungus (and, for that matter, the root) is known.
Until recently, two laborious methods could be used to establish the identity of a fungus on a root tip. The first is to isolate the fungus from a root tip, grow it in the lab, and then introduce it to a tree seedling of the appropriate species and see if mycorrhizas develop. If so, the appearance and structure of the cultivated mycorrhizas are compared to those of the wild mycorrhizas to see whether they are the same. Although this seems simple in concept, its use has been limited by our inability to discover the secret(s) to maintaining most ectomycorrhizal fungi in culture.
The second approach involves tracing the physical connection (hyphae or, more commonly, rhizomorphs) from mushroom to mycorrhizal root tip. To appreciate the difficulty in doing this, next time you're out foraying, give it a try. Start at the base of a Russula, Lactarius, Amanita, or other ectomycorrhizal mushroom and see how far you can trace its mycelium below ground. I suspect you'll quickly develop an appreciation of why morphotyping has proceeded slowly.
Given the amount of time needed to fully develop morphotype descriptions and relate them to specific fungi, it is hardly surprising that most studies that use morphotyping take an abbreviated approach. Whether these less comprehensive approaches will prove sufficient for general use remains to be seen. However, the lack of consistent names for morphotypes might make that concern moot. In most studies using morphotypes, the researchers coin informal names for them, such as "smooth white," "golden yellow," and "black fuzzy." Unfortunately this makes it very difficult to compare different studies because many researchers coin their own names without trying to relate their types to those already described in the literature. Although a scheme for standardizing interim names has been proposed by Reinhard Agerer, it hasn't been widely adopted because of the large amount of work needed to document and publish a standardized name.
All things considered, it isn't surprising that despite the fairly large number of studies of morphotypes that have been conducted, in total they have had a less than overwhelming impact on our knowledge of ectomycorrhizal fungus communities. Additional methods from molecular biology were required before substantial progress could be made.
Use of DNA and other large biochemicals
A variety of different types of complex biochemicals (such as proteins and fatty acids) can now be used to characterize and identify organisms; however, DNA (deoxyribonucleic acid) is the most widely used and will be the focus here. Although the details of the structure and behavior of DNA are fairly complicated (for instance understanding how human DNA that is roughly 2 meters in length can be packaged into 46 chromosomes, each only about 5 micrometers in length, about the same as a small mushroom spore), its basic structure is conceptually simple (Figure 2).
 |
Figure 2: The basic structure of DNA. A DNA molecule can be thought of as an extraordinarily long spiral staircase (a "double helix"), with each leg made up of a string of alternating sugar (S) and phosphate (P) components. To each sugar is attached one of four nucleotides (less formally called bases) -- adenine (A), thymine (T), cytosine (C), or guanine (G). The steps of the staircase, which hold the two legs together, are formed by relatively weak hydrogen bonding between pairs of bases. A and T can bond together, as can C and G, but no other pairs can form. Thus, if you know the sequence of bases in one leg (say, CATCG), you also know what the sequence in the other leg must be (GTAGC). The size or amount of DNA is often referred to in terms of numbers of base-pairs, the number of steps in the staircase. All of the DNA in an organism together is termed its genome. A human genome is comprised of roughly 3 billion base-pairs, while that of a typical fungus is on the order of tens to hundreds of millions of base-pairs. In molecular identification of ectomycorrhizal fungi, either by RFLP or sequencing, DNA segments of perhaps a thousand or so base-pairs are utilized, roughly 0.001 to 0.01% of the fungus's total DNA. |
The structure of DNA has been known since the early 1950's, yet it wasn't until the late 1980's that the stage was set for DNA technology to become commonplace. The key technological advance was development of a procedure called PCR, for polymerase chain reaction. Why was this such a big deal? Recall that most of the DNA of eukaryotic organisms is contained in chromosomes within the nuclei of their cells. Take all the DNA from all the chromosomes and you still don't have much, even if you could be 100% efficient in your extraction. Also keep in mind that we are usually only interested in a very tiny portion of the total DNA. Thus what was needed was a means for making copies of the DNA extracted from an organism; conceptually something like an office photocopier.
That's what PCR does. It allows you to take a very tiny amount of DNA, say from an irreplaceable mushroom type collection, an ancient Egyptian mummy, a 40,000-year-old woolly mammoth, a 30 million-year-old plant fossil, or a blood drop at a crime scene, and by adding DNA building blocks and DNA-building enzyme (DNA polymerase) and subjecting the whole brew to repeated cycles of heating and cooling to produce millions of copies of the bit of DNA you started with. Now you have a sufficient quantity to do things with. Given the huge impact that PCR has had, it's hardly a surprise that the man who conceived it, Kary Mullis, received a Nobel Prize and no doubt now is financially secure, as PCR machines are commonplace in laboratories throughout the world (it's worth noting that the idea for it struck him as he was driving from the San Francisco Bay area to coastal northern California for a weekend getaway -- a good argument for thinking on the highway, instead of talking on your wireless phone or watching DVD's!).
Here are the basics of using DNA to characterize an ectomycorrhiza community. First you collect samples of soil (usually by extracting small cylindrical cores), hopefully ones containing abundant live ectomycorrhizal root tips. You carefully remove the root tips from the soil (a reasonably easy task if your soil is a sandy one, but much more difficult if it is rich in clays), sort them by morphotype (usually using far fewer descriptive characters than a serious morphotyper would), count them, and extract DNA from each tip or small group of tips of the same morphotype. Then, you pop the DNA into your PCR machine, add other necessary ingredients, program the machine, turn it on, let it run overnight, and in the morning, assuming all has worked properly, you have millions of copies of just the bit of DNA you want (only a selected small, much less than 1%, portion of the total DNA of the organism).
All in all, not much harder than using an automatic bread-maker, as long as you can afford the machine, and ingredients that are a bit pricier than flour and yeast. Now that you have a large quantity of DNA, there are several different techniques that can be applied. The two most common ones for mycorrhiza work are fragmentation and sequencing.
Both methods depend on the fact that no two organisms have exactly the same sequence of bases in their DNA, and that the more distant the relationship between two organisms, the more different the base composition of their DNA will be (but note that these differences often are quite small. For instance, the DNA of humans and chimps (our closest relatives) differ by only 1.6%, and that of red-eyed and white-eyed vireos (two species of bird) by just 2.9%.
Sequencing is just what its name suggests -- using rather laborious methods or new (pricey) automatic equipment, you determine the sequence of A's, T's, C's, and G's in one strand of your bit of DNA. Fragmentation involves using so-called "restriction" enzymes to cut the DNA at particular places, such that a collection of fragments of different sizes results. Organisms with different sequences will produce different sets of fragments (these are called restriction fragment length polymorphisms, or RFLP's [pronounced "riff lips"] for short).
The differences are detected by placing the fragmented DNA onto a gel plate that is subjected to an electrical field. Because the DNA fragments carry a net negative charge, they migrate in a line through pores in the gel toward the positive-charged end of the plate. Small fragments move faster, large fragments move slower, and through comparison with fragments of known size, the size of each of the sample fragments can be determined. The cover of Mushroom the Journal, Issue #50 (Winter 1995-96) shows the fragment patterns from several morels. Samples from the same, or closely related, species should yield the same pattern of fragment sizes.
But what can you tell from a sequence such as AATCGCTTACCGG, or a picture of fragment bands on a gel? Nothing directly -- you need to find a match with a sequence or fragment pattern from a known entity, such as a confidently identified mushroom. Thus, collecting and identifying mushrooms or other fungus parts is an essential step in building a reference library of sequences or fragment patterns (increasingly these libraries exist as computer databases, accessible to researchers via the internet). If you find that the sequence or pattern from your unknown mycorrhizal root tip matches the sequence or pattern from a known mushroom, then you can conclude that that is the fungus in your mycorrhiza (or at least a very closely related one).
What have we learned using these methods?
Two recent review articles provide excellent summaries of the state of ectomycorrhiza community studies. One is by a Swede, Anders Dahlberg, and the other by two American Toms -- Tom Horton of Syracuse University and Tom Bruns of UC Berkeley (references are provided at the end of this article). The two reviews tell much the same story.
Mushrooms Provide only a Partial Picture
The bottom line is that the pictures we get from above-ground studies (mushroom collecting) are not the same as, and can be greatly different than, those we get using below-ground methods, either morphotyping or molecular.
Although there were earlier indications, the first emphatic demonstration of the non-concurrence of the above-ground and below-ground views was provided by Monique Gardes and Tom Bruns in a study of a Bishop pine (Pinus muricata) woodland north of San Francisco Bay. The most abundant mushrooms in that forest by a wide margin are Suillus pungens and Amanita franchetii. Yet the Suillus was found in only small amounts on roots, and the Amanita in but moderate amounts. Each species often failed to be found on roots collected immediately beneath a mushroom of that species.
 |
Figure 3: Mushroom surveys provide only a partial, and possibly misleading, picture of the ectomycorrhiza community. Thanks to David Read for the ostrich idea, and especially to Sam Norris for converting my rough sketches and scribbling into elegant artwork. |
Overall, the roots were dominated by a Russula, R. amoenolens, a resupinate fungus, Tomentella sublilacina, and a few other taxa. Many others were present, but not abundant. Only one mushroom of R. amoenolens was found during the entire 3-year study, and the Tomentella was overlooked for much of it. Thus, the below-ground community was very different than the mushrooms suggested it would be. DNA data from a number of subsequent studies, mostly in western North America and northern Europe, have generally confirmed the pattern observed by Gardes and Bruns.
Thus, fungus species that produce readily observable mushrooms make up a minority (perhaps only 20 to 30%) of the root inhabitants, and so mushroom surveys alone cannot provide us with more than a partial picture of the ectomycorrhizal fungus community (Figure 3). Among the mushroom and truffle-formers, species of Russula, Lactarius, Suillus, and their hypogeous relatives, such as Rhizopogon, typically are most abundant.
So, what are the other 70 to 80%? Some are species that don't seem to produce fruiting bodies at all or, if they do, not often enough or in sufficient quantity for us to find them. Cenococcum graniforme is an example. Others, such as species of Tomentella, produce inconspicuous resupinate (paint-smear) fruiting bodies (Volume 2 of Fungi of Switzerland has a good collection of photos of these macroscopically less than thrilling fungi). Generally we have assumed that resupinate fungi are all wood-rotters, so it has been a surprise to find that many of them actually are mycorrhizal.
Forests Have Many Ectomycorrhizal Fungi
Even though only tiny areas are actually covered by the soil samples, large numbers of taxa can be found. For instance, in 14 studies reviewed by Horton and Bruns, as many as 135 taxa were found in aggregate sampling areas of less than one-half square meter, a small door mat at most. This richness in small areas is not surprising if you consider that a boreal spruce-pine forest can have 10,000 to 100,000 or more root tips in each square meter of forest floor, approximately equal to the number of needles directly overhead. Thus, ectomycorrhiza communities are rich in species. However, just as in the Toft and Orton study with mushrooms, the DNA data show that only a few taxa are common, most are rare or at most uncommon.
Are Things the Same Everywhere?
Just as it's rare to find a single species of mushroom spread evenly over a large area of forest -- instead we tend to find one here and a few there, scattered about in small, or sometimes large, groups -- ectomycorrhizas tend to be distributed in patchy fashion. They are present and maybe even abundant in some spots, but absent elsewhere. For instance, in an Arizona woodland composed of scattered pinyon pine trees, Catherine Gehring and her co-workers found that the root systems of individual trees were dominated by a small number of fungus species and that different trees were dominated by different fungi.
Another example of the patchy distribution of ectomycorrhizal fungi is provided by the association between snow plant (Sarcodes sanguinea, a non-chlorophyll-containing plant related to Indian pipe and pine drops) and the false truffle Rhizopogon ellenae in red fir (Abies magnifica) forests in the Sierra Nevada. Martin Bidartondo, Annette Kretzer, and their colleagues found that R. ellenae made up 86-98% of the red-fir ectomycorrhizas associated with the snow plant root masses, but that its abundance declined rapidly until they were absent just one-half meter from each snow plant. A second species of Rhizopogon also occurred in the study plots, but not in the areas occupied by the snow plants.
The extent to which the communities change with time (even over the course of a year) is not known but, given the patchiness of their spatial distributions, seems likely to be significant. For instance, here in the Pacific Northwest there are mushrooms, such as Cortinarius violaceus that are only found in old forests, whereas other species seem to exhibit preferences for younger forests (see the three-chanterelle discussion in my review of the MSA meeting on page 20 of the last issue). Are these patterns related to changes that occur on the roots, or do they reflect peculiarities of mushroom fruiting? Either way, it could have important implications for how the forests function.
What about other Mycorrhiza Types?
The impetus for most of the work done until recently came from the relative ease of compiling DNA profile libraries for mushrooms (even though this is no simple task, and so only a small proportion of all mushroom species have been profiled thus far). Because arbuscular mycorrhizal fungi and ericoid mycorrhizal fungi produce no fruiting bodies or, at most, small inconspicuous ones, studies of these communities have been far fewer than those of ectomycorrhizal fungi. However, as the molecular methods have become better understood and applied in more sophisticated ways, studies of arbuscular mycorrhiza and ericoid mycorrhiza communities are becoming much more common. It is still too early to draw many generalizations from them, however. (Studies of AM fungi generally have relied on spores recovered from soils. For a fascinating discussion of the results of some of these studies, see the cover story of the November 2001 BioScience.)
Cautionary Notes
Very Few Types of Forest Have Been Studied
Investigation of ectomycorrhizal fungus communities is still in its infancy. Thus, we must take care not to embrace too many conclusions too strongly. For example, most studies have been conducted in relatively simple forests of pines (e.g., Scots, jack, and Bishop pines) and/or spruce (Norway and Sitka spruce), so it would be premature to assume that what we've learned will apply everywhere. As a wider variety of habitats is investigated, it will be interesting to see whether the current picture extends to forests with other ectomycorrhizal trees such as Douglas-fir, hemlocks, firs, oaks, beech, eucalyptus, and dipterocarps, especially where they occur in complex mixtures of species.
Our Understanding of Basic Mycorrhizal Biology Is Limited
There's also still a lot of basic biology we don't understand, so we need to keep our minds open as we proceed. For instance at ICOM-2, colleagues of Reinhard Agerer reported that, immediately beneath a mushroom of Russula exalbicans growing under birch trees, they found a mycorrhizal root tip that had the morphological and anatomical characteristics of the ectomycorrhizas formed by Lactarius pubescens. No russula-type mycorrhizas were found. RFLP analysis of the fungus DNA extracted from the root tip showed a combined pattern of the Lactarius and the Russula. Thus, two fungi were present within one apparently homogeneous mycorrhiza. The same phenomenon was found in a different set of birch, Russula, and Lactarius species. We know little about how common this might be, and even less about what it might mean physiologically and ecologically. What other surprises might be waiting out there?
Perhaps the existence of dual-fungus mycorrhizas will help explain why many studies have had such low rates of success in matching root-tip species with mushrooms. For instance, a study in Sweden found 135 taxa based on RFLP patterns from root tips, and 66 species of ectomycorrhizal mushrooms. However, only 11 matches were found between the two groups. Although two-in-one mycorrhizas might be one factor contributing to this problem, chances are that it isn't the major one.
What about the Hyphae in the Soil?
Advances in molecular technology have allowed us to expand our inquiries from mushrooms to root tips. However, there still is the extraradical mycelium to wonder about -- the hyphae spread throughout the soil that are doing the mycorrhizal grunt work. As such, it represents an essential part of the physiological and ecological functions of ectomycorrhizas. Efforts currently are underway to develop methods for identifying and quantifying the extraradical community, taking many cues from bacterial ecologists. However, at this point, we know nearly nothing about it. If I had to predict what will transpire eventually, it would be that a third picture will appear, different from those produced by mushrooms and root tips.
Wrap-up
Through mushroom surveys, morphotyping, and molecular studies, we are beginning to develop a picture of the communities of ectomycorrhizal fungi in some of our temperate forests and woodlands. The most productive approach involves combined use of all three methods.
Although rapid progress is being made, we still have much to learn, especially about habitats that have received little or no attention, and about the extraradical phase of the fungi. As this sort of knowledge accumulates and we fill in our rosters of players, it will be important to shift focus to physiological studies of the different fungi and fungus-plant combinations. These hold the key to answering the interesting fundamental questions such as "why are there so many fungi?"
Next time
In Installment 5, we'll look at mycorrhizal networks, groups of plants interconnected by fungal hyphae into a "wood-wide web." What is the evidence for their existence, and what might their ecological implications be?
References
(* = review article: these contain references to many of the individual studies that have been carried out)
- Agerer, Reinhard. 1987 et seq. Colour Atlas of Ectomycorrhizae. Einhorn-Verlag, Schwabisch Gmund, Munich.
- *Bever, James D., Peggy A. Schultz, Anne Pringle, and Joseph B. Morton. 2001. Arbuscular mycorrhizal fungi: more diverse than meets the eye, and the ecological tale of why. BioScience 51(11): 923-931 (November).
- *Dahlberg, Anders. 2001. Community ecology of ectomycorrhizal fungi: an advancing interdisciplinary field. New Phytologist 150: 555-562.
- Gardes, M. and T.D. Bruns. 1996. Community structure of ectomycorrhizal fungi in a Pinus muricata forest: above- and below-ground views. Canadian Journal of Botany 74: 1572-1583.
- Goodman, D.M., D.M. Durall, J.A. Trofymow, and S.M. Berch. 1996 et seq. A Manual of Concise Descriptions of North American Ectomycorrhizae. Mycologue Publications. Sidney, B.C., Canada.
- *Horton, Thomas R., and Thomas D. Bruns. 2001. The molecular revolution in ectomycorrhizal ecology: peeking into the black-box. Molecular Ecology 10: 1855-1871.
- Dimensions: e.g., length of system, length and diameter of individual root tips
- Branching pattern: e.g., dichotomous (wish-bone, or tuning fork), pinnate (like a fern frond; see Figure 1f), coralloid (see Figure 1e)
- Texture of mantle surface: e.g., smooth, matte, grainy, fuzzy
- Color and color changes
- Chemical reactions: e.g., with potassium hydroxide (KOH) or Melzer's reagent
- Anatomy of mantle: e.g., arrangement and shape of cells and hyphae
- Mantle layers: e.g., number, type, and thickness of layers
- Cell lengths and widths
- Hyphae: e.g., diameter, ornamentation, presence/density of emanating hyphae
- Anatomy of mycelial strands (rhizomorphs)
- Cystidia: e.g., presence, shape, encrustations
- Chlamydospores: e.g., presence, shape, size
- UV-fluorescence: e.g., color, intensity
Copyright Notice
This material appeared in slightly modified form in Mushroom: the Journal of Wild Mushrooming, Issue 75, Spring 2002. It can be used freely for not-for-profit personal and educational purposes provided that its source is clearly credited.

